Forensic Genomics: Genomic discovery of forensically relevant variation as revealed through massively parallel sequencing
Improvements in sequencing technology have resulted in the ability to simultaneously generate detailed genetic information across a large number of genetic loci from a single biological sample in a forensic context. This work explores sequence variation of forensically relevant STR and SNP loci, and includes the international STRseq project (https://www.ncbi.nlm.nih.gov/bioproject/380127).
Traditionally, DNA profiling in a forensic genetics context has relied on a class of DNA marker called short tandem repeats (STRs), where variation is assessed through examining the difference in length of the DNA in individuals, at the specific genomic locations where these markers are present. Improvements in sequencing technology with the commercial availability of massively parallel sequencing (MPS) means that we can now analyse the actual genetic sequences at these locations rather than only assess the length of the sequence, and this raises the possibility that we can now detect variation by both sequence and length at the same time. This project aims to sequence DNA from over 1000 UK resident individuals in 5 population groups using the Verogen DNA Signature Prep kit which simultaneously amplifies 27 autosomal STRs and 94 autosomal single nucleotide polymorphism markers (SNPs). Sequence variation at these markers can then be catalogued and investigated: an important advance both to determine how useful this MPS technology may be to forensic genetics and to provide sequence allele frequency data that can be subsequently used for other purposes. Detailed analysis of the data can highlight where useful sequence variation is observed – in which markers, and in the repeat or flanking regions – as well as determine the extent to which sequence variation is population-specific, and hence whether this type of data can be used for ancestry analysis (i.e. can it be used to also determine what region of the world an individual’s ancestor may be from, whether that be, for example, Europe, Africa, or East Asia).
We aim to improve forensic inference from such compromised samples by adding sequence information to standard typed variants and the local chromosomal environment to improve the capability to make use of such data in an international setting.
Aims
- To investigate the sequence variation present within 27 autosomal forensic STR markers in 5 major populations groups along with 94 identity SNPs and provide population specific frequencies
- To identify the value of both repeat and flanking sequence variation associated with these STR and SNP markers
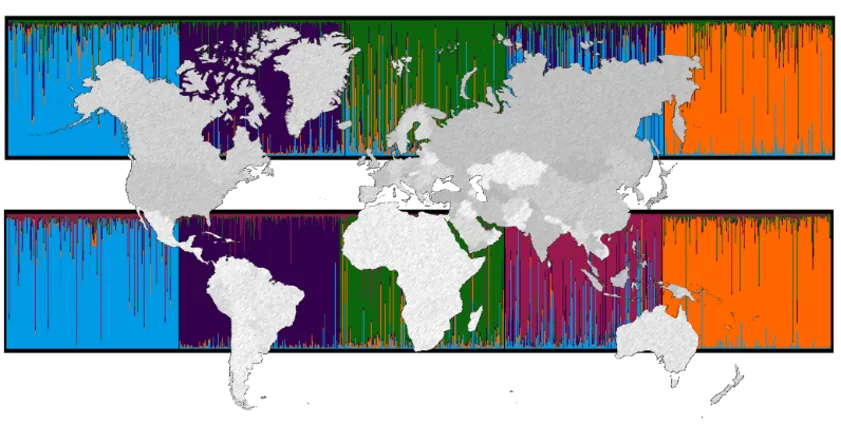
- To explore the utility of this variation for ancestry analysis from a forensic STR profile
Our Partners

University of Santiago de Compostela

University of Innsbruck

Verogen
Principal Investigator
Investigators
Affiliations
Funding
Funding Body: Royal Commission For The Exhibition of 1851
Amount: £28,000
Period: September 2017 - October 2022






