What are non equilibrium systems?
How do we control heating in fast electronic devices, or understand how next-generation solar cells might work? Can insights into extreme events in the sciences be used to detect whether a network of financial institutions is close to a crash? Can we predict the fate of cellular states in health and disease? How do we understand whether epidemics will spread, or predict the likelihood of extreme meteorological events? How do we optimize the design of nanoparticles to maximize efficiency in drug and gene delivery?
These research challenges all relate to non-equilibrium systems. Such systems are typically irreversible, e.g. because they dissipate energy persistently pumped in from the outside, or they age towards an equilibrium state that is never reached. They also frequently exhibit extreme events, such as cascades of collective failure or the thermally activated rare event of a drug molecule being released from a vesicle. In contrast to phenomena at, or close to, equilibrium, our understanding of non-equilibrium systems is still in its infancy.
As the examples above show, this challenge is central to a wide variety of problems across the spectrum of physical, mathematical, biological and environmental sciences. Significant advances will have to come from researchers capable of exploiting and enhancing these interdisciplinary links.
Overview of Centre
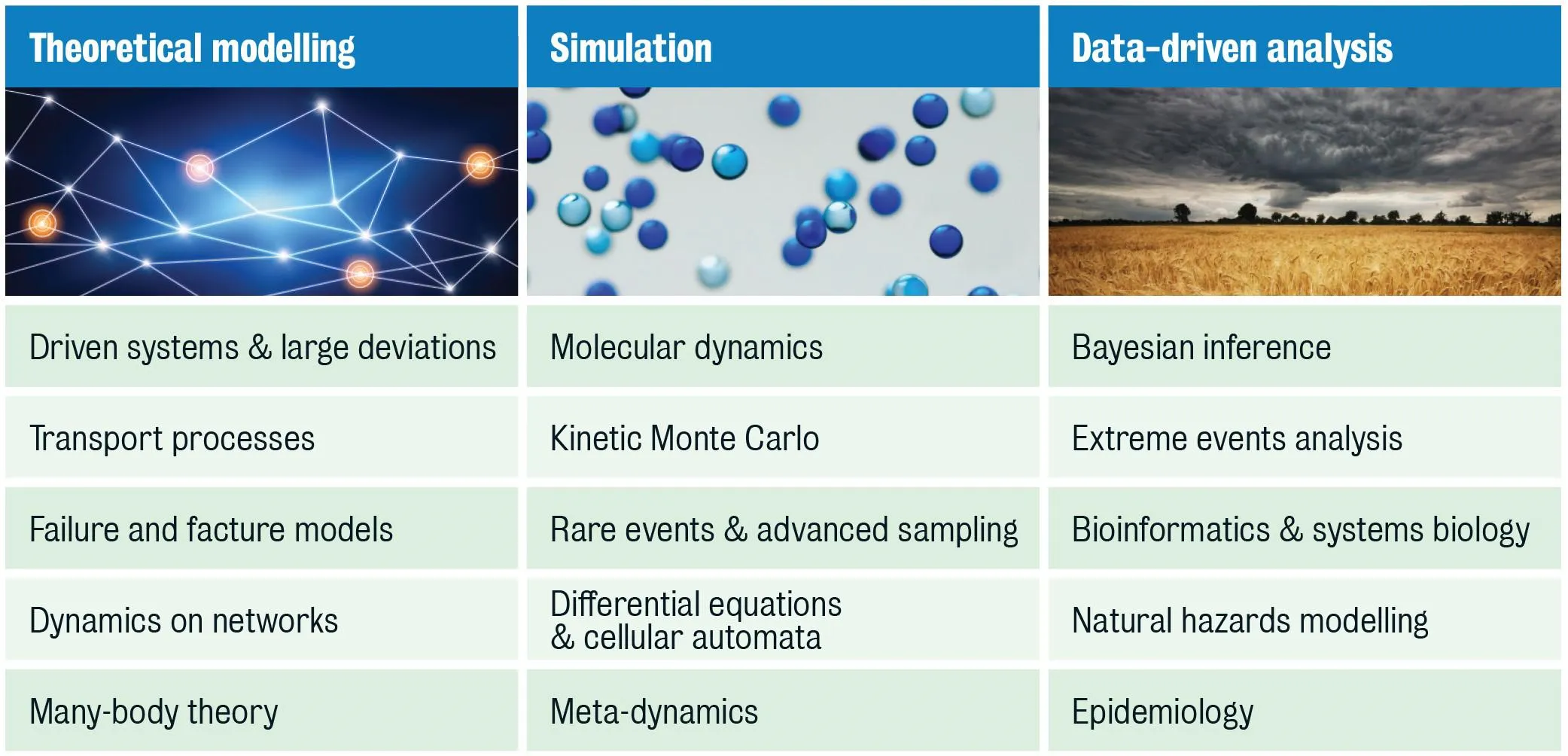
Non-equilibrium processes underpin many challenging problems across the natural sciences. The objectives of CANES are to train a new generation of researchers in cross-disciplinary approaches to non-equilibrium systems and develop deeper insights into non-equilibrium processes focusing on the three key strands of theoretical modelling, simulation and data-driven analysis.
The mission of CANES is to train future research leaders in the understanding, control and design of systems far from equilibrium, based on rigorous training in theoretical modelling, simulation and data-driven analysis, and a breadth of awareness of common themes across disciplines. Additionally, CANES will also function as a UK Centre of Excellence for the research and research user community, and a national and international hub in the area of non-equilibrium systems.
Studentships and Eligibility
The ultimate goal is to address interdisciplinary challenges:
- How do we characterize, design and grow materials, and devices, with novel properties out of equilibrium?
- How do we control and exploit the stochastic processes inherent to biological systems?
- Can we use inference and information assimilation approaches from physics and biology to monitor and evaluate the state and direction of non-equilibrium environmental systems?
CANES draws on a broad range of supervisor expertise in Mathematics, Physics, Chemistry, Informatics, Computational and Systems Biomedicine, Earth and Environmental Sciences, including partners at Imperial College London, University College London and Queen Mary London.
Each year the centre will offer 10 fully-funded 4-year PhD studentships at King's College London. CANES Studentships covers course fees, a stipend for living expenses (ca £16,000 per year), and conference travel and internship funds.
The programme can support UK applicants as well as a limited number of students from the EU and overseas.
Funding for CANES students covers course fees, a stipend for living expenses (ca £16,000 per year), and conference travel and internship funds.
(Please note CANES CDT is currently not accepting new applications for the programme)
Training programme: year 1
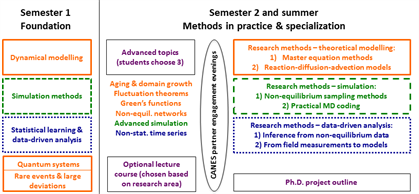
Semester 1
CANES students will attend 4 lecture courses in the first semester; for the fourth module they can choose between two options, with a classical or quantum focus, depending on their research interests:
- 7CCMNE01 Dynamical Modelling of Non-Equilibrium Systems
- 7CCPNE02 Simulation Methods for Non-Equilibrium Systems
- 7CCMNE03 Statistical Learning and Data-Driven Analysis
- 7CCPNE05 Modelling Quantum Many-body Systems (Option 1)
- 7CCMNE06 Rare Events and Large Deviations (Option 2)
The first 2-3 weeks of semester 1 will have an additional booster weeks programme to level up any differences in background preparation of a new CANES cohort. The cohort tutor will provide help and advice with module selection.
Semester 2
In the second semester students will attend an “Advanced topics in non-equilibrium systems” and an optional lecture course chosen from the wide range of available modules according to research interest.
Three Research Methods modules will occupy the rest of the second semester and the summer. These will be closely aligned with the first semester lecture blocks on theoretical modelling, data-driven analysis, and simulation, in order to integrate taught material and hands-on research training. Each research methods module will consist of a mini-project, which will be assessed by a project report and presentation.
- 7CCMNE07 Research Methods for Theoretical Modelling of Non-Equilibrium Systems
- 7CCMNE08 Research Methods for Simulation of Non-Equilibrium Systems
- 7CCMNE09 Research Methods for Data-Driven Analysis of Non-Equilibrium Systems
- 7CCMNE04 Advanced Topics in Non-Equilibrium Systems, 15 credits
- optional Module
Training programme: years 2-4
After year 1, emphasis will shift to research and away from assessed courses. Structured training will be delivered by a number of routes including:
● Open questions sandpits where students and leading experts meet for 2 days to identify the most urgent questions and promising areas for progress in a given field
● A broad spectrum of focussed master classes
● A weekly cross-CDT seminar series
● A journal club where students present papers in pairs to encourage peer-to-peer learning
● An annual retreat to foster research interactions, along with sessions on ideas generation, creative problem solving techniques, and working in multi-disciplinary teams offered by an experienced trainer
● A two-day residential course (launch pad workshop from the Engentia Open programme) with a focus on encouraging and developing innovative collaborative projects, across discipline and sector boundaries. This offers researchers a unique opportunity to understand cross-disciplinary collaboration right at the beginning of their career.
● Transferable skills training through the KCL Graduate School
● Practical training in outreach and public understanding of science via contributions to existing outreach channels like taster days and teachers' conferences
● Dedicated careers sessions delivered by King's College Careers Service
Internships:
To ensure that CANES students are exposed to a range of research environments, the Centre will organize the opportunity for one internship of ca. two months' duration for each student.
We have secured agreement to host internships from a number of industrial partners as well as overseas academic institutions, including:
- National Physical Laboratory
- The Francis Crick Institute
- Fios Genomics
- Financial Network Analytics
- Microsoft Research
- Universität Potsdam (Germany)
- Technische Universität Berlin (Germany)
- Forschungszentrum Jülich (Germany)
- Université Pierre et Marie Curie, Paris (France)
- Université Montpellier 2 (France)
- University of California, Irvine (USA)
- Wuhan University (China)
Eligibility:
General entry advice
- The programme can support UK applicants as well as a limited number of students from the EU and overseas.
- Bachelor's degree with first class honours (or international equivalent) in any relevant discipline including mathematics, physics, chemistry, engineering, materials science, biophysics, geophysical sciences and computer science.
- Note: A 2:1 (upper second class) honours degree (or international equivalent) may be acceptable depending on the candidate's academic background.
- A strong background in quantitative science will be essential for students to take full advantage of the CANES training programme.
English language requirements for International Students
To follow this programme successfully, you must have a good command of English and be able to apply this in an academic environment. Therefore, you are usually required to provide certificated proof of your competence in English before starting your studies.
To apply to the CANES programme you will need to meet the minimum English Language Requirements please as specified here. For the CANES programme you will need to meet the requirements for the Faculty of Natural & Mathematical Sciences as detailed for each of the English Language Tests.
Our Partners

The Francis Crick Institute

National Physics Laboratory

AWE

Citibank

Deutsche Bank

Deutsche Zentral-Genossenschaftbank

Financial Network Analytics

Fios Genomics

Forwardlane

Lucid

Microsoft Research Cambridge

ONRG

Seldon

The Simons Foundation

Techstars
Contact us
CANES CDT
Faculty of Natural & Mathematical Sciences King’s College London
Strand, London
WC2R 2LS
Be sure to check and like our Facebook page!
You can also find us on twitter: @canes_cdt
