When we started to work with high dimensional IMC data a couple of years ago, there were not many tools for their analysis. It has been a lot of fun do develop our own tool thanks to the talent of Michele Bortolomeazzi, a PhD student funded by the CRUK KHP Centre , and taking advantage of the vast expertise in tissue histology of Professor Jo Spencer.
Professor Francesca Ciccarelli
09 February 2022
Kings researchers develop a new software for the analysis of multiplexed tissue imaging data
A team of researchers from the Schools of Cancer & Pharmaceutical Sciences and Immunology and Microbial Sciences and the Cancer Systems Biology laboratory at the Francis Crick Institute, led by Professor Francesca Ciccarelli, have published new paper showcasing novel software for the analysis of complex tissue imaging data.
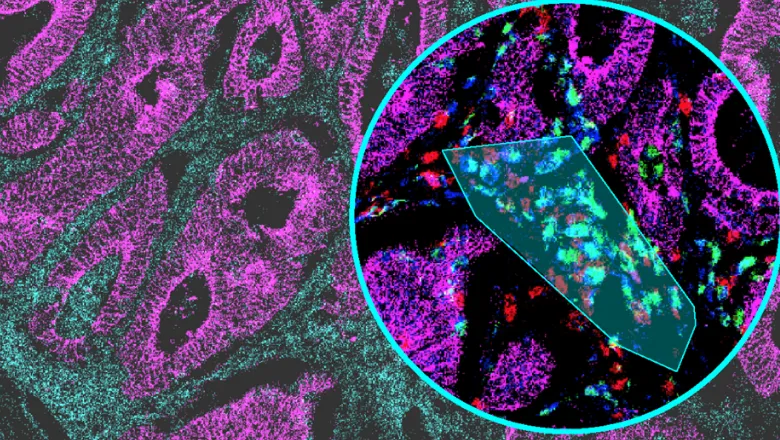
Recently developed multiplexed imaging technologies, such imaging mass cytometry (IMC), enable detailed investigation of tissue cell composition that are of particular value for mapping the tissue-level characteristics of disease and predicting the outcome of therapies that depend on the tissue environment, such as cancer immunotherapy. These techniques usually produce large amounts of high dimensional data that are difficult to analyse and often require multiple tools and users with computational expertise.
SIMPLI (Single-cell Identification from MultiPlexed Images) is a flexible software that unifies all steps of multiplexed imaging data analysis, providing an improved alternative to high-dimension imaging data analysis also for non-specialists. SIMPLI performs a spatially resolved, single-cell analysis of each tissue slide as well as cell-independent quantifications of marker expression to investigate features undetectable at the cell level. It also produces tabular and graphical outputs that faciliate the visual inspection of results at each step of the analysis.
To test the performance and versatility of SIMPLI, the researchers analysed images that had been obtained with different imaging technologies, including but not limited to IMC, and were diverse in origin, size and resolution.
King’s College London has been one of the first Institutions in Europe to host a Helios imaging mass cytometer, which is available in the BRC Flow Cytometry Platform.
SIMPLI is freely accessible at https://github.com/ciccalab/SIMPLI.
The study describing SIMPLI is published in Nature Communications.


